Visually communicate your scientific data with R
Designing scientific figures
ARNA, INSERM U1212, CNRS UMR 5320, Université de Bordeaux
UFR des Sciences Pharmaceutiques, Université de Bordeaux
April 22, 2026
Introduction to Grammar of Graphics
A framework for data visualization
Introduced by Leland Wilkinson (The Grammar of Graphics, 2005).
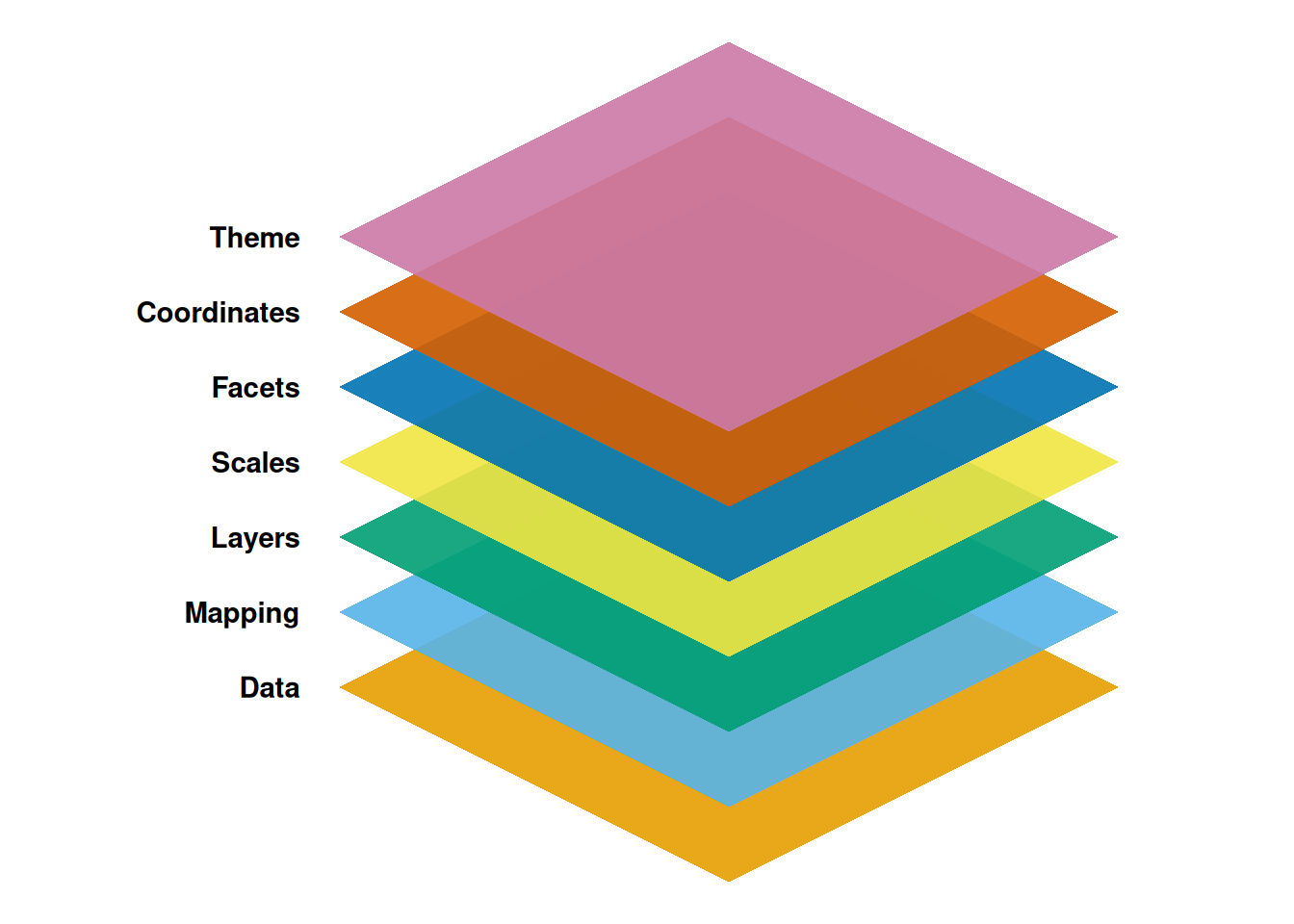
Graphics can be built up from a set of components:
- Data: the dataset to be visualized
- Variables: mapping of objects to values
- Algebra: operations to combine variables and specify dimensions
- Scales: represent variables on measured dimensions
- Statistics: functions to change the appearance and representation
- Geometry: creation of geometric graphs from variables
- Coordinates: mapping to coordinate systems
- Aesthetics: sensory attributes
- Facets: subplots based on subsets of data
- Guides: legends and axes to explain the graph
Adapted to R in the ggplot2 package
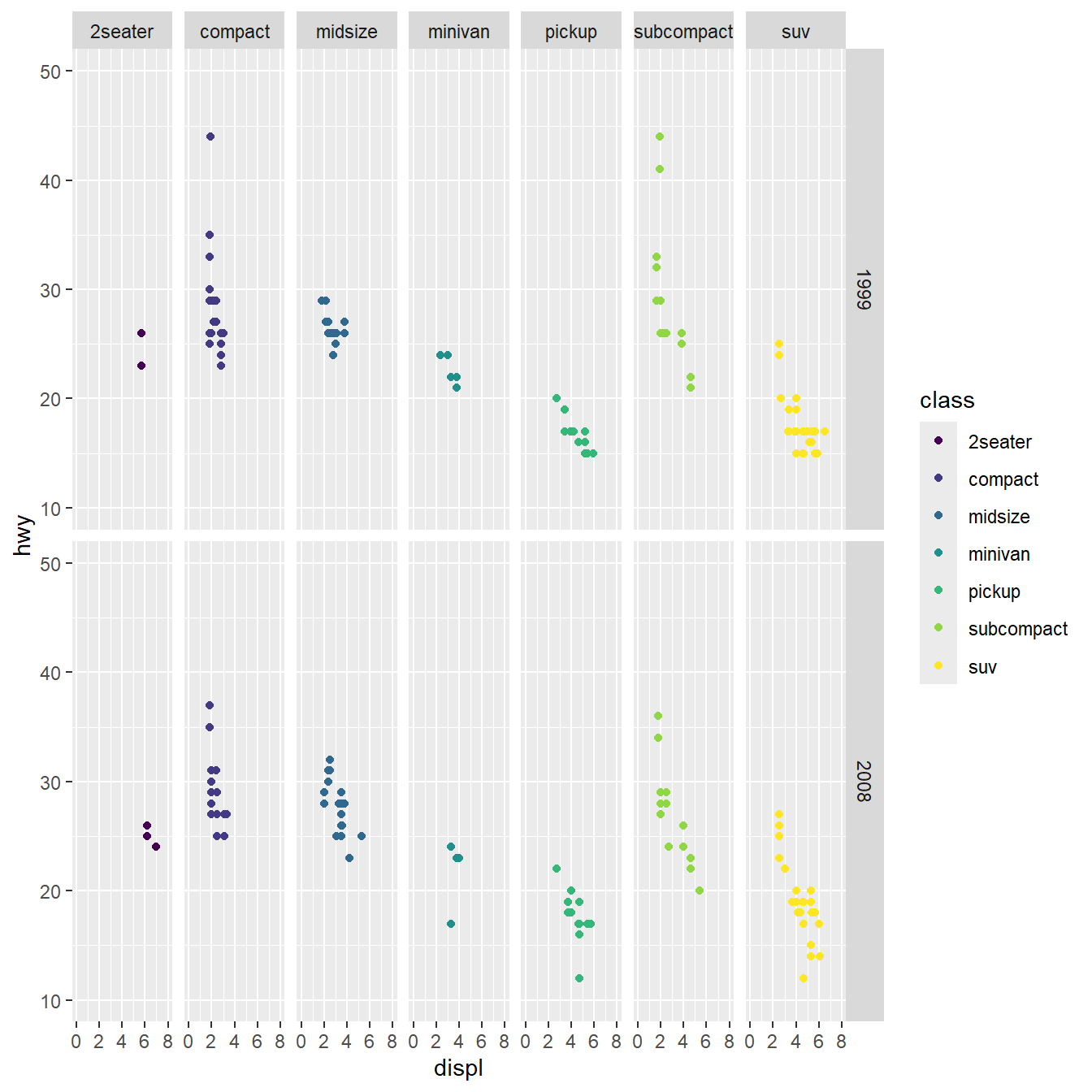
Example: Data should be formatted with 1 column per variable
ggplot initialization provides a plotting area
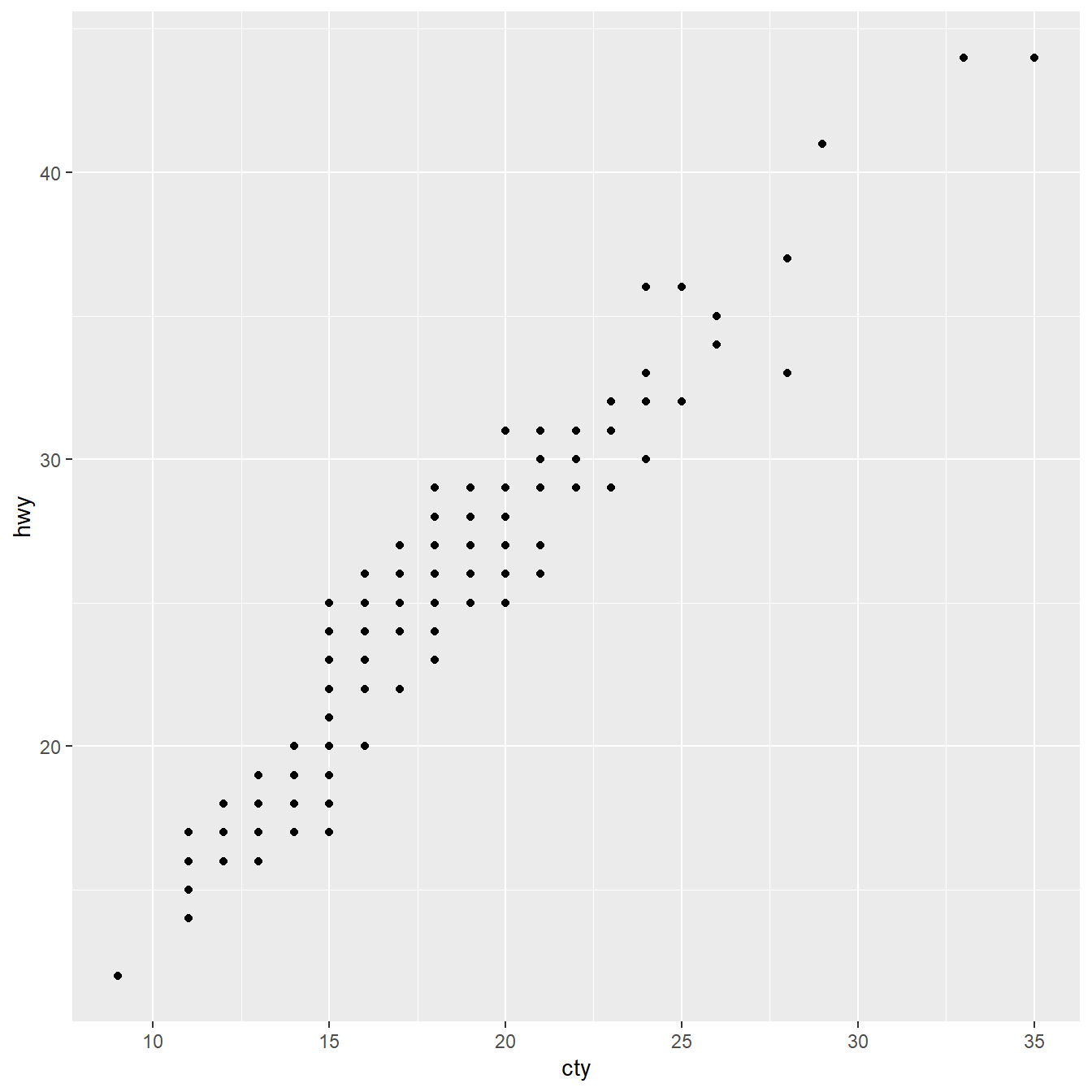
Mapping x and y coordinates provide axes
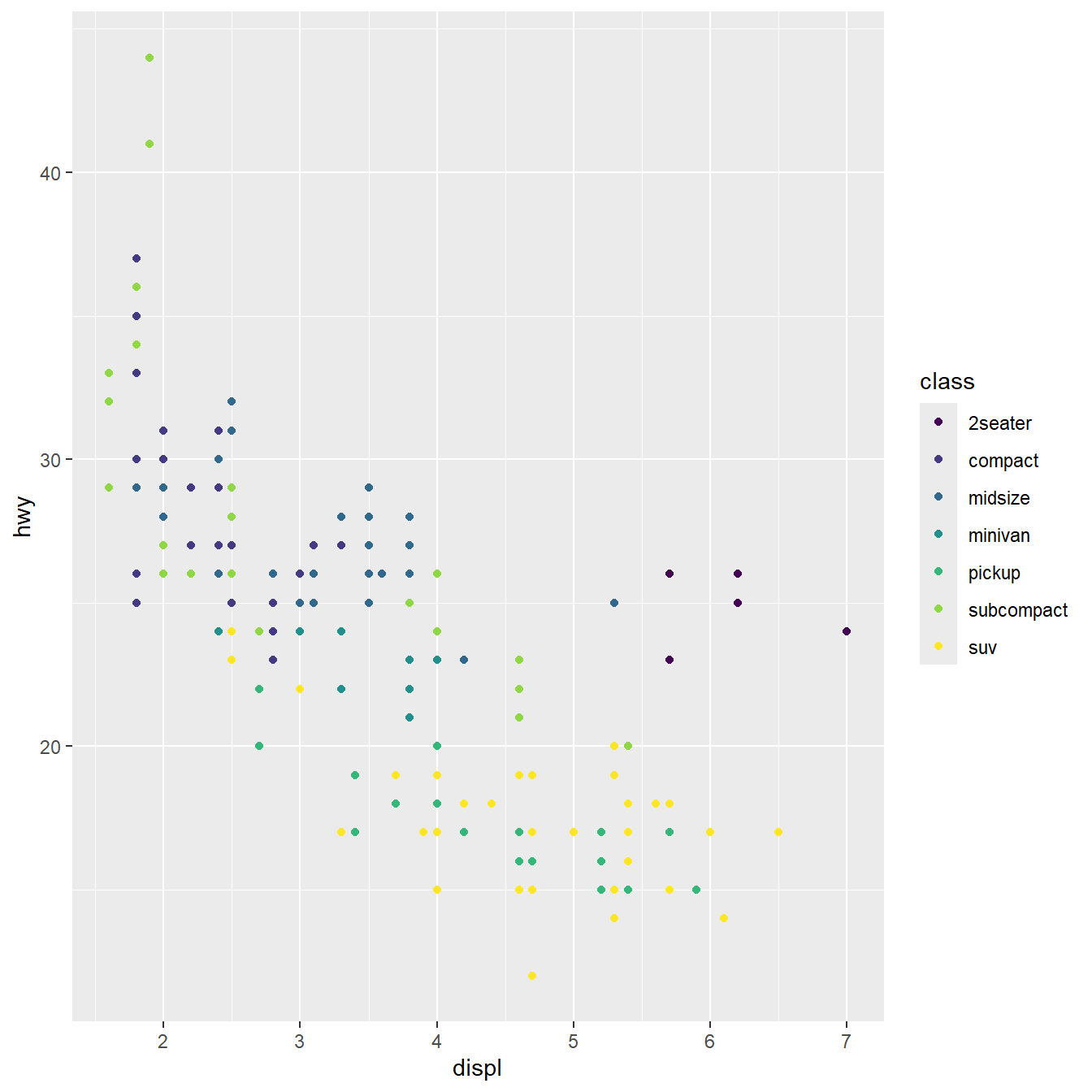
Geometries can be added as layers
Scales can be applied to aesthetics
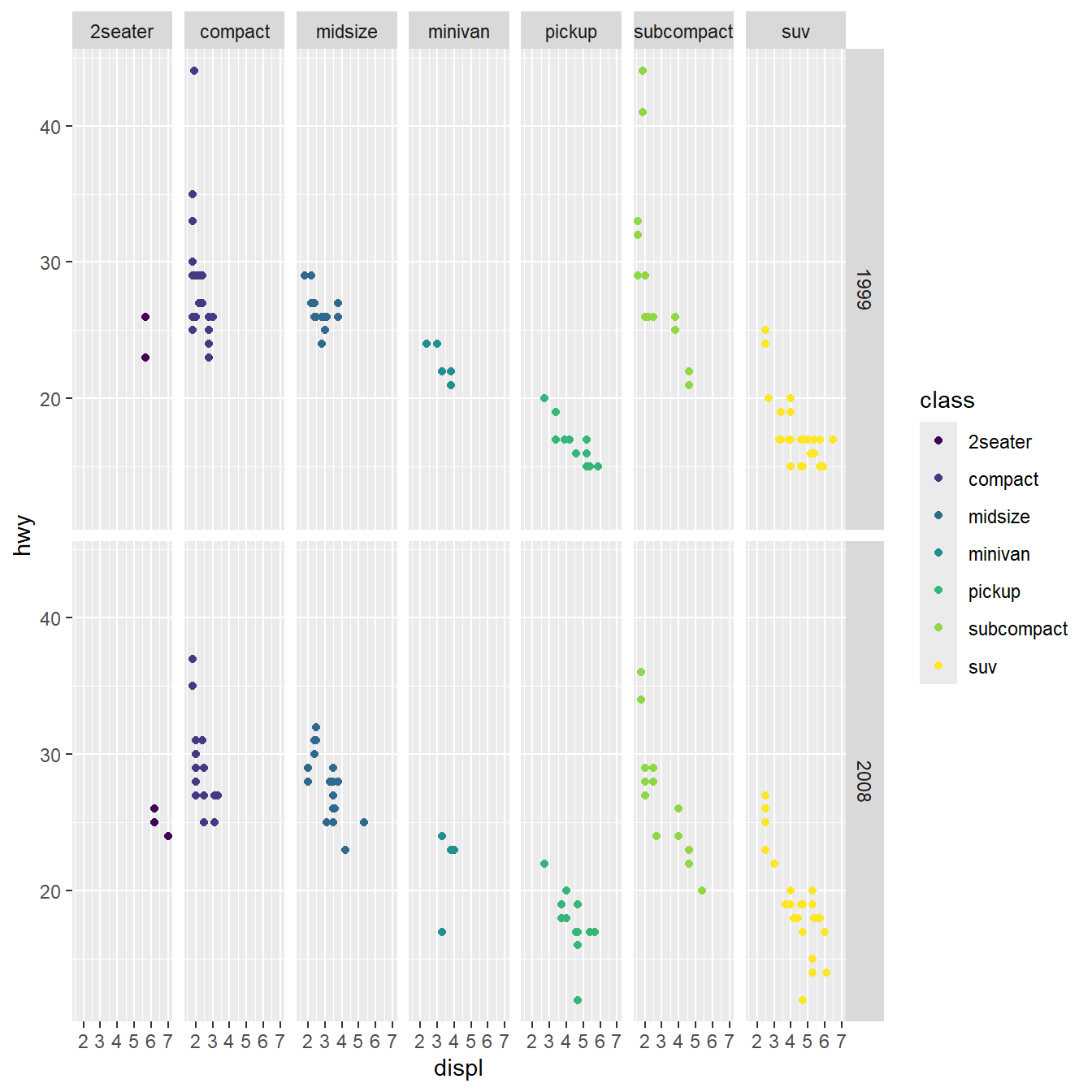
Facetting allows generating many subplots
Coordinates allows re-scaling
Make it pretty/corporate/legible with a theme
library(ggplot2)
ggplot(
mpg,
mapping = aes(
x = displ,
y = hwy,
colour = class
)
) +
geom_point() +
scale_colour_viridis_d() +
facet_grid(year ~ class) +
coord_cartesian(
xlim = c(0, 8),
ylim = c(10, 50),
clip = "off"
) +
theme_minimal() +
theme(
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
legend.position = "bottom",
axis.title = element_text(size = 14, face = "bold"),
axis.line = element_line(color = "black", size = 0.75),
axis.ticks = element_line(color = "black", size = 0.75),
legend.title = element_text(face = "bold", size = 10),
legend.text = element_text(size = 10),
strip.text = element_text(face = "bold", size = 10)
) +
scale_x_continuous(
name = "Engine Displacement (L)",
breaks = seq(0, 8, 4),
limits = c(0, 8),
expand = c(0, 0)
) +
scale_y_continuous(name = "Highway Miles per Gallon") +
labs(colour = "Vehicle Class")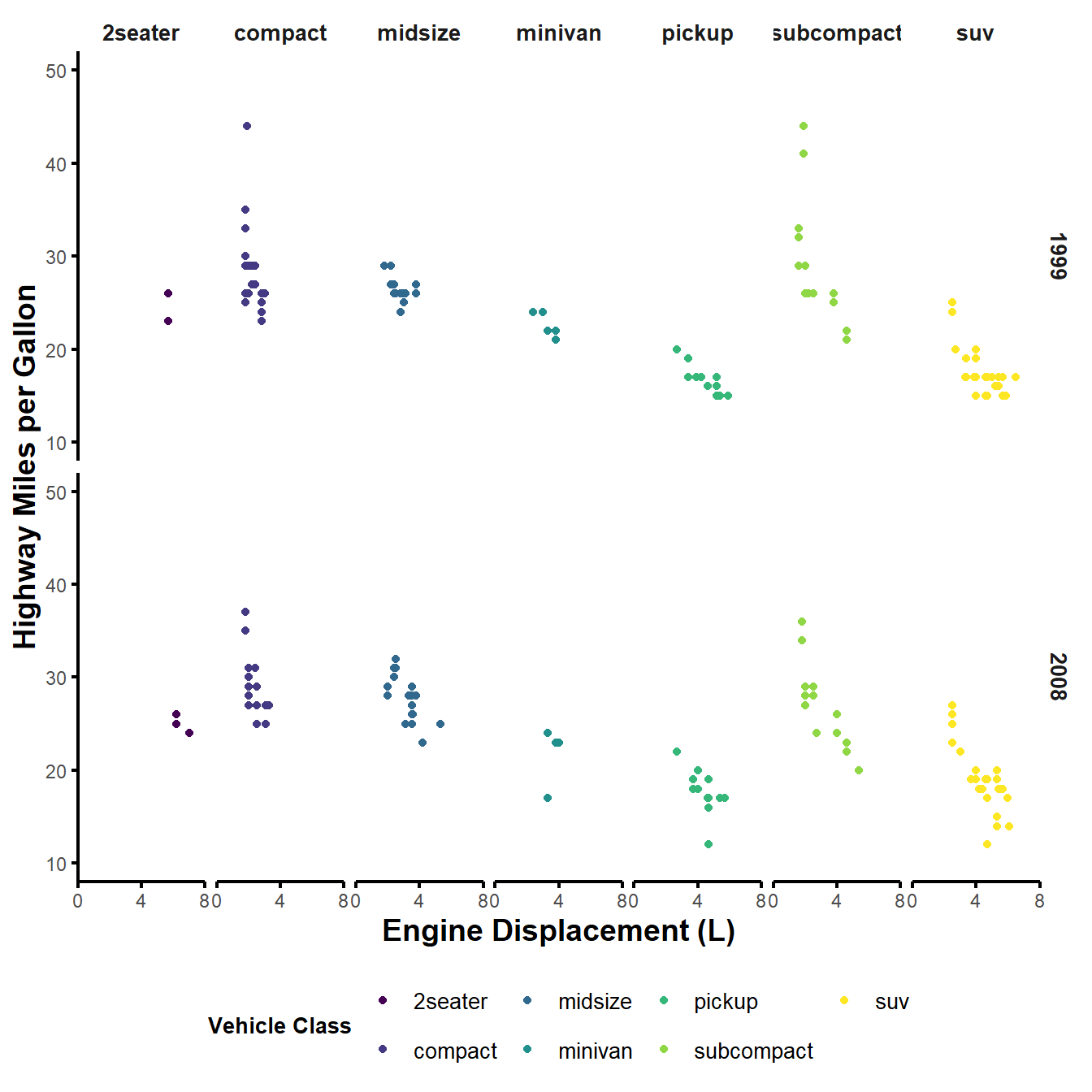
Setup of the environment
Tidyverse install verification
Data.table install verification
Reminders on variables and data types and modes
Assigning and printing numeric variables
Assigning and printing character variables
Assigning and printing boolean variables
Null, missing values and infinities
a_null <- NULL
a_missing <- NA
a_nan <- NaN
a_inf <- Inf
a_minus_inf <- -Inf
char_missing <- NA_character_
real_missing <- NA_real_
int_missing <- NA_integer_
str(
list(
a_null = a_null,
a_missing = a_missing,
a_nan = a_nan,
a_inf = a_inf,
a_minus_inf = a_minus_inf,
char_missing = char_missing,
real_missing = real_missing,
int_missing = int_missing
)
)List of 8
$ a_null : NULL
$ a_missing : logi NA
$ a_nan : num NaN
$ a_inf : num Inf
$ a_minus_inf : num -Inf
$ char_missing: chr NA
$ real_missing: num NA
$ int_missing : int NAA NULL value is not missing
a_null <- NULL
a_missing <- NA
a_nan <- NaN
a_inf <- Inf
a_minus_inf <- -Inf
char_missing <- NA_character_
real_missing <- NA_real_
int_missing <- NA_integer_
cat(
"NULL represents the absence of a value or object,",
"\nwhile NA represents a missing value in a vector.",
"\n\nThe length of a NULL value is",
length(a_null),
",\nwhile the length of a missing value is",
length(a_missing),
"."
)NULL represents the absence of a value or object,
while NA represents a missing value in a vector.
The length of a NULL value is 0 ,
while the length of a missing value is 1 .Factors are important for ordering elements in visualizations
[1] low medium high medium low
Levels: high low mediumCreating vectors
[1] "1" "2" "3" "4" "5" "6" "7" "8"
[9] "9" "10" "coucou"R coerces all elements to the most flexible type
Subsetting vectors by position, logical and negative indexing
[1] "1" "2" "3" "coucou"Naming vector elements
Accessing named vector elements with [] and [[]]
Modifying elements and types of a vector
[1] "one" "2" "42" "4" "5" Creating and inspecting lists
# lists cannot contain
# different types of data
# but can contain different types of objects,
# including vectors
a_list <- list(
numeric_vector = c(1, 2, 3, 4),
character_vector = c("a", "b", "c"),
mixed_vector = c(1, "b", TRUE),
nested_list = list(
numeric_vector = c(5, 6, 7, 8),
character_vector = c("d", "e", "f"),
mixed_vector = c(9, "g", FALSE)
)
)
a_list$numeric_vector
[1] 1 2 3 4
$character_vector
[1] "a" "b" "c"
$mixed_vector
[1] "1" "b" "TRUE"
$nested_list
$nested_list$numeric_vector
[1] 5 6 7 8
$nested_list$character_vector
[1] "d" "e" "f"
$nested_list$mixed_vector
[1] "9" "g" "FALSE"Accessing list elements by name with $
Accessing nested list elements by name with $
Comparing list elements for equality
Subsetting lists: [ ] vs. [[ ]]
a_list <- list(
numeric_vector = c(1, 2, 3, 4),
character_vector = c("a", "b", "c"),
mixed_vector = c(1, "b", TRUE),
nested_list = list(
numeric_vector = c(5, 6, 7, 8),
character_vector = c("d", "e", "f"),
mixed_vector = c(9, "g", FALSE)
)
)
str(
list(
'dollar_name' = a_list$character_vector,
'bracket_name' = a_list[["character_vector"]],
'bracket_index' = a_list[[2]],
'bracket_twice' = a_list[2][[1]]
)
)List of 4
$ dollar_name : chr [1:3] "a" "b" "c"
$ bracket_name : chr [1:3] "a" "b" "c"
$ bracket_index: chr [1:3] "a" "b" "c"
$ bracket_twice: chr [1:3] "a" "b" "c"What happens if we access a_list[2] vs. a_list[[2]]?
Creating matrices
Indexing matrices by row and column
Creating dataframes as named list of vectors
Accessing column names with names()
Accessing dataframe columns with $
[1] "A" "A" "B" "B" "C"Accessing dataframe columns with $
[1] "A" "B" "C"Subsetting dataframes by row/column indices
id group value
1 1 A 10.2
3 3 B 9.8Subsetting dataframes by column names
id value
1 1 10.2
2 2 12.5
3 3 9.8
4 4 11.1
5 5 13.0Subsetting dataframes by logical conditions
id group value
1 1 A 10.2
2 2 A 12.5
4 4 B 11.1
5 5 C 13.0Subsetting dataframes can be subsetted with subset()
Finding row indices with which()
[1] 1 2Evaluating expressions in datafames with with()
tibble::tibble() is a modern dataframe
Tibbles have improved defaults
'data.frame': 5 obs. of 3 variables:
$ id : int 1 2 3 4 5
$ group: chr "A" "A" "B" "B" ...
$ value: num 10.2 12.5 9.8 11.1 13tibble [5 × 3] (S3: tbl_df/tbl/data.frame)
$ id : int [1:5] 1 2 3 4 5
$ group: chr [1:5] "A" "A" "B" "B" ...
$ value: num [1:5] 10.2 12.5 9.8 11.1 13 id group value
1 1 A 10.2
2 2 A 12.5
3 3 B 9.8
4 4 B 11.1
5 5 C 13.0Subsetting with dplyr::filter() and dplyr::select()
# A tibble: 3 × 3
id group value
<int> <chr> <dbl>
1 1 A 10.2
2 2 A 12.5
3 4 B 11.1Dataframes and tibbles can be subsetted with dplyr functions
id value
1 1 10.2
2 2 12.5
3 3 9.8
4 4 11.1
5 5 13.0The data.table alternative for large dataset and concise syntax
Think in terms of basic units: rows, columns, and groups.
DT[i, j, by]
- i: row selection (filtering)
- j: column selection (projection)
- by: grouping operations
data.table is optimized for speed and memory, ideal for large datasets
The data.table alternative for large dataset and concise syntax
id group value
<int> <char> <num>
1: 1 A 10.2
2: 2 A 12.5
3: 3 B 9.8
4: 4 B 11.1
5: 5 C 13.010 common pitfalls
And how to avoid them
R is 1-indexed
[1] "a"[ ] returns a listis not [[ ]]
Check for missing values with is.na()
R coerces all elements to the most flexible type
Modifying a subset does not modify the original
[1] "a" "b"R recycles shorter vectors to match the length of longer vectors
NULL is different from NA
list_NA_NULL <- list(
"missing" = NA,
"null_value" = NULL
)
list_NA_NULL["length_missing"] <- length(list_NA_NULL[[1]])
list_NA_NULL["length_null"] <- length(list_NA_NULL[[2]])
list_NA_NULL[5] <- length(list_NA_NULL[2])
list_NA_NULL["na_check"] <- is.na(list_NA_NULL["missing"])
list_NA_NULL["null_check"] <- is.null(list_NA_NULL[["null_value"]])
list_NA_NULL["null_check_2"] <- is.null(list_NA_NULL["null_value"])
# length_null = 0 because NULL has no length
# [5] = 1 because list_NA_NULL[2] is a list of length 1, not NULL itself
# null_check = TRUE because the element inside is NULL
# null_check_2 = FALSE because it is a list of length 1 != NULL
str(list_NA_NULL)List of 8
$ missing : logi NA
$ null_value : NULL
$ length_missing: int 1
$ length_null : int 0
$ : int 1
$ na_check : logi TRUE
$ null_check : logi TRUE
$ null_check_2 : logi FALSEAdding a factor level does not add data
[1] A B C
Levels: A B C D[[]] allow dynamic df access, $ does not
[1] 1 2 3 4 5Use drop = FALSE to retain df structure when subsetting a single column
# A tibble: 5 × 2
id even
<dbl> <dbl>
1 1 2
2 2 4
3 3 6
4 4 8
5 5 10Basic operations
Basic operations
Basic operations
Basic operations
Reminders on basic functions
Basic functions
Basic functions
Basic functions
Data preparation
Starting dataset
# A tibble: 6 × 11
manufacturer model displ year cyl trans drv cty hwy fl class
<chr> <chr> <dbl> <int> <int> <chr> <chr> <int> <int> <chr> <chr>
1 audi a4 1.8 1999 4 auto(l5) f 18 29 p compa…
2 audi a4 1.8 1999 4 manual(m5) f 21 29 p compa…
3 audi a4 2 2008 4 manual(m6) f 20 31 p compa…
4 audi a4 2 2008 4 auto(av) f 21 30 p compa…
5 audi a4 2.8 1999 6 auto(l5) f 16 26 p compa…
6 audi a4 2.8 1999 6 manual(m5) f 18 26 p compa…Use of glimpse
Rows: 234
Columns: 11
$ manufacturer <chr> "audi", "audi", "audi", "audi", "audi", "audi", "audi", "…
$ model <chr> "a4", "a4", "a4", "a4", "a4", "a4", "a4", "a4 quattro", "…
$ displ <dbl> 1.8, 1.8, 2.0, 2.0, 2.8, 2.8, 3.1, 1.8, 1.8, 2.0, 2.0, 2.…
$ year <int> 1999, 1999, 2008, 2008, 1999, 1999, 2008, 1999, 1999, 200…
$ cyl <int> 4, 4, 4, 4, 6, 6, 6, 4, 4, 4, 4, 6, 6, 6, 6, 6, 6, 8, 8, …
$ trans <chr> "auto(l5)", "manual(m5)", "manual(m6)", "auto(av)", "auto…
$ drv <chr> "f", "f", "f", "f", "f", "f", "f", "4", "4", "4", "4", "4…
$ cty <int> 18, 21, 20, 21, 16, 18, 18, 18, 16, 20, 19, 15, 17, 17, 1…
$ hwy <int> 29, 29, 31, 30, 26, 26, 27, 26, 25, 28, 27, 25, 25, 25, 2…
$ fl <chr> "p", "p", "p", "p", "p", "p", "p", "p", "p", "p", "p", "p…
$ class <chr> "compact", "compact", "compact", "compact", "compact", "c…Use of select
Rows: 234
Columns: 4
$ displ <dbl> 1.8, 1.8, 2.0, 2.0, 2.8, 2.8, 3.1, 1.8, 1.8, 2.0, 2.0, 2.8, 2.8,…
$ hwy <int> 29, 29, 31, 30, 26, 26, 27, 26, 25, 28, 27, 25, 25, 25, 25, 24, …
$ class <chr> "compact", "compact", "compact", "compact", "compact", "compact"…
$ year <int> 1999, 1999, 2008, 2008, 1999, 1999, 2008, 1999, 1999, 2008, 2008…